Transcription Factors as Early Diagnostic Biomarkersfor Chronic Kidney Disease: A Comprehensive Analysis
DOI:
https://doi.org/10.71321/1pe9g186Keywords:
Chronic kidney disease, Transcription factor, Biomarker, Data analysisAbstract
Background: Chronic kidney disease (CKD) is a global health concern with significant implications for public health and mortality rates, projected to become the fifth leading cause of death by 2040. The search for early diagnostic targets for CKD is imperative. In this study, we concentrated on identifying key transcription factors (TFs) for the early diagnosis of CKD and established a regulatory network between these TFs and their corresponding target genes.
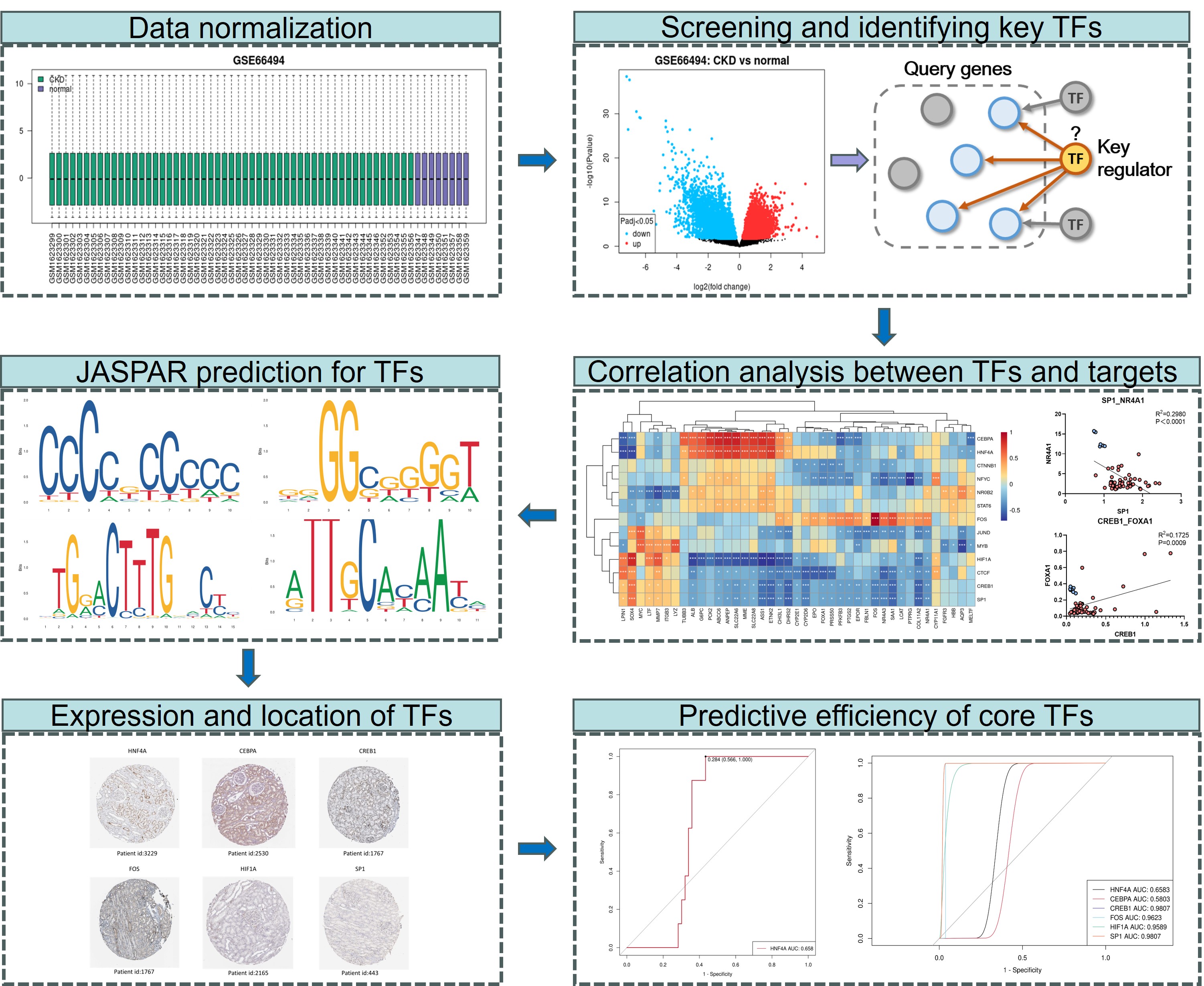
Methods: We conducted microarray data analysis and Gene Set Enrichment Analysis (GSEA) to identify differentially expressed genes (DEGs) and the associated pathways in CKD. We further explored the potential regulatory TFs among DEGs using the TRRUST v2 database and validated the TF-target regulatory relationships through correlation analysis and the JASPAR database. The protein expression of the identified TFs in renal tissues was also assessed.
Results: The analysis identified six TFs, namely HNF4A, CEBPA, CREB1, FOS, HIF1A, and SP1, which demonstrated potential as diagnostic biomarkers for CKD. These TFs showed differentially expressed patterns in CKD and were found to have multiple regulatory relationships with DEGs, indicating their crucial role in the disease process. ROC analysis revealed high predictive efficiency for four of these TFs (CREB1, FOS, HIF1A, and SP1), while the combined predictive efficiency of all TFs was exceptionally high.
Conclusion: Our findings highlight the role of transcription factors in the pathophysiological process of CKD and identify several key TFs with potential for clinical translation as early diagnostic biomarkers for the disease. Further validation and exploration are warranted to leverage the potential clinical utility of these TFs in the early diagnosis and prognosis of CKD.
References
[1] Webster AC, Nagler EV, Morton RL, & Masson P. (2017). Chronic Kidney Disease. Lancet, 389(10075), 1238-1252. https://doi.org/10.1016/S0140-6736(16)32064-5
[2] Jiang M, Bai M, Lei J, Xie Y, Xu S, Jia Z, et al. (2020). Mitochondrial dysfunction and the AKI-to-CKD transition. Am J Physiol Renal Physiol, 319(6), F1105-F1116. https://doi.org/10.1152/ajprenal.00285.2020
[3] Chen TK, Knicely DH, & Grams ME. (2019). Chronic Kidney Disease Diagnosis and Management: A Review. JAMA, 322(13), 1294-1304. https://doi.org/10.1001/jama.2019.14745
[4] Chawla LS, Eggers PW, Star RA, & Kimmel PL. (2014). Acute kidney injury and chronic kidney disease as interconnected syndromes. N Engl J Med, 371(1), 58-66. https://doi.org/10.1056/NEJMra1214243
[5] Tepus M, Tonoli E, & Verderio EAM. (2022). Molecular profiling of urinary extracellular vesicles in chronic kidney disease and renal fibrosis. Front Pharmacol, 13, 1041327. https://doi.org/10.3389/fphar.2022.1041327
[6] Humphreys BD. (2018). Mechanisms of Renal Fibrosis. Annu Rev Physiol, 80, 309-326. https://doi.org/10.1146/annurev-physiol-022516-034227
[7] Tang PM, Nikolic-Paterson DJ, & Lan HY. (2019). Macrophages: versatile players in renal inflammation and fibrosis. Nat Rev Nephrol, 15(3), 144-158. https://doi.org/10.1038/s41581-019-0110-2
[8] Li L, Fu H, & Liu Y. (2022). The fibrogenic niche in kidney fibrosis: components and mechanisms. Nat Rev Nephrol, 18(9), 545-557. https://doi.org/10.1038/s41581-022-00590-z
[9] Chen S, Liu YH, Dai DP, Zhu ZB, Dai Y, Wu ZM, et al. (2021). Using circulating O-sulfotyrosine in the differential diagnosis of acute kidney injury and chronic kidney disease. BMC Nephrol, 22(1), 66. https://doi.org/10.1186/s12882-021-02268-3
[10] Lambert SA, Jolma A, Campitelli LF, Das PK, Yin Y, Albu M, et al. (2018). The Human Transcription Factors. Cell, 172(4), 650-665. https://doi.org/10.1016/j.cell.2018.01.029
[11] Ruiz-Ortega M, Rayego-Mateos S, Lamas S, Ortiz A, & Rodrigues-Diez RR. (2020). Targeting the progression of chronic kidney disease. Nat Rev Nephrol, 16(5), 269-288. https://doi.org/10.1038/s41581-019-0248-y
[12] Hishikawa A, Hayashi K, & Itoh H. (2018). Transcription Factors as Therapeutic Targets in Chronic Kidney Disease. Molecules, 23(5). https://doi.org/10.3390/molecules23051123
[13] Ma TT, & Meng XM. (2019). TGF-beta/Smad and Renal Fibrosis. Adv Exp Med Biol, 1165, 347-364. https://doi.org/10.1007/978-981-13-8871-2_16
[14] Li W, Feng W, Su X, Luo D, Li Z, Zhou Y, et al. (2022). SIRT6 protects vascular smooth muscle cells from osteogenic transdifferentiation via Runx2 in chronic kidney disease. J Clin Invest, 132(1). https://doi.org/10.1172/JCI150051
[15] Yang Y, Yu X, Zhang Y, Ding G, Zhu C, Huang S, et al. (2018). Hypoxia-inducible factor prolyl hydroxylase inhibitor roxadustat (FG-4592) protects against cisplatin-induced acute kidney injury. Clin Sci (Lond), 132(7), 825-838. https://doi.org/10.1042/CS20171625
[16] Wen Y, Lu X, Ren J, Privratsky JR, Yang B, Rudemiller NP, et al. (2019). KLF4 in Macrophages Attenuates TNFalpha-Mediated Kidney Injury and Fibrosis. J Am Soc Nephrol, 30(10), 1925-1938. https://doi.org/10.1681/ASN.2019020111
[17] Huang Y, Wang S, Zhou J, Liu Y, Du C, Yang K, et al. (2020). IRF1-mediated downregulation of PGC1alpha contributes to cardiorenal syndrome type 4. Nat Commun, 11(1), 4664. https://doi.org/10.1038/s41467-020-18519-0
[18] Tang TT, Wang B, Li ZL, Wen Y, Feng ST, Wu M, et al. (2021). Kim-1 Targeted Extracellular Vesicles: A New Therapeutic Platform for RNAi to Treat AKI. J Am Soc Nephrol, 32(10), 2467-2483. https://doi.org/10.1681/ASN.2020111561
[19] Wang J, Ge S, Wang Y, Liu Y, Qiu L, Li J, et al. (2021). Puerarin Alleviates UUO-Induced Inflammation and Fibrosis by Regulating the NF-kappaB P65/STAT3 and TGFbeta1/Smads Signaling Pathways. Drug Des Devel Ther, 15, 3697-3708. https://doi.org/10.2147/DDDT.S321879
[20] Nakagawa S, Nishihara K, Miyata H, Shinke H, Tomita E, Kajiwara M, et al. (2015). Molecular Markers of Tubulointerstitial Fibrosis and Tubular Cell Damage in Patients with Chronic Kidney Disease. PLoS One, 10(8), e0136994. https://doi.org/10.1371/journal.pone.0136994
[21] Lyu F, Han F, Ge C, Mao W, Chen L, Hu H, et al. (2023). OmicStudio: A composable bioinformatics cloud platform with real-time feedback that can generate high-quality graphs for publication. Imeta, 2(1), e85. https://doi.org/10.1002/imt2.85
[22] Han H, Cho JW, Lee S, Yun A, Kim H, Bae D, et al. (2018). TRRUST v2: an expanded reference database of human and mouse transcriptional regulatory interactions. Nucleic Acids Res, 46(D1), D380-D386. https://doi.org/10.1093/nar/gkx1013
[23] Zhou W, Sha Y, Zeng J, Zhang X, Zhang A, & Ge X. (2021). Computational Systems Pharmacology, Molecular Docking and Experiments Reveal the Protective Mechanism of Li-Da-Qian Mixture in the Treatment of Glomerulonephritis. J Inflamm Res, 14, 6939-6958. https://doi.org/10.2147/JIR.S338055
[24] Castro-Mondragon JA, Riudavets-Puig R, Rauluseviciute I, Lemma RB, Turchi L, Blanc-Mathieu R, et al. (2022). JASPAR 2022: the 9th release of the open-access database of transcription factor binding profiles. Nucleic Acids Res, 50(D1), D165-D173. https://doi.org/10.1093/nar/gkab1113
[25] Yu X, Xu M, Meng X, Li S, Liu Q, Bai M, et al. (2020). Nuclear receptor PXR targets AKR1B7 to protect mitochondrial metabolism and renal function in AKI. Sci Transl Med, 12(543). https://doi.org/10.1126/scitranslmed.aay7591
[26] Ming WH, Luan ZL, Yao Y, Liu HC, Hu SY, Du CX, et al. (2023). Pregnane X receptor activation alleviates renal fibrosis in mice via interacting with p53 and inhibiting the Wnt7a/beta-catenin signaling. Acta Pharmacol Sin, 44(10), 2075-2090. https://doi.org/10.1038/s41401-023-01113-7
[27] You R, Zhou W, Li Y, Zhang Y, Huang S, Jia Z, et al. (2020). Inhibition of ROCK2 alleviates renal fibrosis and the metabolic disorders in the proximal tubular epithelial cells. Clin Sci (Lond), 134(12), 1357-1376. https://doi.org/10.1042/CS20200030
[28] Romagnani P, Remuzzi G, Glassock R, Levin A, Jager KJ, Tonelli M, et al. (2017). Chronic kidney disease. Nat Rev Dis Primers, 3, 17088. https://doi.org/10.1038/nrdp.2017.88
[29] Edeling M, Ragi G, Huang S, Pavenstadt H, & Susztak K. (2016). Developmental signalling pathways in renal fibrosis: the roles of Notch, Wnt and Hedgehog. Nat Rev Nephrol, 12(7), 426-439. https://doi.org/10.1038/nrneph.2016.54
[30] Schunk SJ, Floege J, Fliser D, & Speer T. (2021). WNT-beta-catenin signalling - a versatile player in kidney injury and repair. Nat Rev Nephrol, 17(3), 172-184. https://doi.org/10.1038/s41581-020-00343-w
[31] Zhang Y, Ding X, Guo L, Zhong Y, Xie J, Xu Y, et al. (2023). Comprehensive analysis of the relationship between xanthine oxidoreductase activity and chronic kidney disease. iScience, 26(11), 107332. https://doi.org/10.1016/j.isci.2023.107332
[32] Saucedo AL, Perales-Quintana MM, Paniagua-Vega D, Sanchez-Martinez C, Cordero-Perez P, & Minsky NW. (2018). Chronic Kidney Disease and the Search for New Biomarkers for Early Diagnosis. Curr Med Chem, 25(31), 3719-3747. https://doi.org/10.2174/0929867325666180307110908
[33] Muto Y, Wilson PC, Ledru N, Wu H, Dimke H, Waikar SS, et al. (2021). Single cell transcriptional and chromatin accessibility profiling redefine cellular heterogeneity in the adult human kidney. Nat Commun, 12(1), 2190. https://doi.org/10.1038/s41467-021-22368-w
[34] Marable SS, Chung E, Adam M, Potter SS, & Park JS. (2018). Hnf4a deletion in the mouse kidney phenocopies Fanconi renotubular syndrome. JCI Insight, 3(14). https://doi.org/10.1172/jci.insight.97497
[35] Hamilton AJ, Bingham C, McDonald TJ, Cook PR, Caswell RC, Weedon MN, et al. (2014). The HNF4A R76W mutation causes atypical dominant Fanconi syndrome in addition to a beta cell phenotype. J Med Genet, 51(3), 165-169. https://doi.org/10.1136/jmedgenet-2013-102066
[36] Wu B, & Brooks JD. (2012). Gene expression changes induced by unilateral ureteral obstruction in mice. J Urol, 188(3), 1033-1041. https://doi.org/10.1016/j.juro.2012.05.004
[37] Kasano-Camones CI, Takizawa M, Ohshima N, Saito C, Iwasaki W, Nakagawa Y, et al. (2023). PPARalpha activation partially drives NAFLD development in liver-specific Hnf4a-null mice. J Biochem, 173(5), 393-411. https://doi.org/10.1093/jb/mvad005
[38] Yang T, Poenisch M, Khanal R, Hu Q, Dai Z, Li R, et al. (2021). Therapeutic HNF4A mRNA attenuates liver fibrosis in a preclinical model. J Hepatol, 75(6), 1420-1433. https://doi.org/10.1016/j.jhep.2021.08.011
[39] Fujiu K, Manabe I, & Nagai R. (2011). Renal collecting duct epithelial cells regulate inflammation in tubulointerstitial damage in mice. J Clin Invest, 121(9), 3425-3441. https://doi.org/10.1172/JCI57582
[40] Gao R, Wu Y, Yang Q, Chen L, Chen J, Wang B, et al. (2023). The Interaction of Apelin and FGFR1 Ameliorated the Kidney Fibrosis through Suppression of TGFbeta-Induced Endothelial-to-Mesenchymal Transition. Oxid Med Cell Longev, 2023, 5012474. https://doi.org/10.1155/2023/5012474
[41] Liu G, Ding W, Neiman J, & Mulder KM. (2006). Requirement of Smad3 and CREB-1 in mediating transforming growth factor-beta (TGF beta) induction of TGF beta 3 secretion. J Biol Chem, 281(40), 29479-29490. https://doi.org/10.1074/jbc.M600579200
[42] Yano N, Suzuki D, Endoh M, Zhang W, Xu YC, Padbury JF, et al. (2012). In vitro silencing of the insulin receptor attenuates cellular accumulation of fibronectin in renal mesangial cells. Cell Commun Signal, 10(1), 29. https://doi.org/10.1186/1478-811X-10-29
[43] Shan Q, Zheng G, Zhu A, Cao L, Lu J, Wu D, et al. (2016). Epigenetic modification of miR-10a regulates renal damage by targeting CREB1 in type 2 diabetes mellitus. Toxicol Appl Pharmacol, 306, 134-143. https://doi.org/10.1016/j.taap.2016.06.010
[44] Silverstein DM, Travis BR, Thornhill BA, Schurr JS, Kolls JK, Leung JC, et al. (2003). Altered expression of immune modulator and structural genes in neonatal unilateral ureteral obstruction. Kidney Int, 64(1), 25-35. https://doi.org/10.1046/j.1523-1755.2003.00067.x
[45] Zhang Y, Feng XH, & Derynck R. (1998). Smad3 and Smad4 cooperate with c-Jun/c-Fos to mediate TGF-beta-induced transcription. Nature, 394(6696), 909-913. https://doi.org/10.1038/29814
[46] Singhal PC, Sharma P, Sanwal V, Prasad A, Kapasi A, Ranjan R, et al. (1998). Morphine modulates proliferation of kidney fibroblasts. Kidney Int, 53(2), 350-357. https://doi.org/10.1046/j.1523-1755.1998.00758.x
[47] Zhao H, Han Y, Jiang N, Li C, Yang M, Xiao Y, et al. (2021). Effects of HIF-1alpha on renal fibrosis in cisplatin-induced chronic kidney disease. Clin Sci (Lond), 135(10), 1273-1288. https://doi.org/10.1042/CS20210061
[48] Li X, Yang S, Yan M, Guan N, Li J, Xie Q, et al. (2020). Interstitial HIF1A induces an estimated glomerular filtration rate decline through potentiating renal fibrosis in diabetic nephropathy. Life Sci, 241, 117109. https://doi.org/10.1016/j.lfs.2019.117109
[49] Li ZL, Wang B, Lv LL, Tang TT, Wen Y, Cao JY, et al. (2021). FIH-1-modulated HIF-1alpha C-TAD promotes acute kidney injury to chronic kidney disease progression via regulating KLF5 signaling. Acta Pharmacol Sin, 42(12), 2106-2119. https://doi.org/10.1038/s41401-021-00617-4
[50] Kassimatis TI, Nomikos A, Giannopoulou I, Lymperopoulos A, Moutzouris DA, Varakis I, et al. (2010). Transcription factor Sp1 expression is upregulated in human glomerulonephritis: correlation with pSmad2/3 and p300 expression and renal injury. Ren Fail, 32(2), 243-253. https://doi.org/10.3109/08860220903411164
[51] Jiang L, Zhou Y, Xiong M, Fang L, Wen P, Cao H, et al. (2013). Sp1 mediates microRNA-29c-regulated type I collagen production in renal tubular epithelial cells. Exp Cell Res, 319(14), 2254-2265. https://doi.org/10.1016/j.yexcr.2013.06.007
[52] Zhou W, Fang J, Jia Q, Meng H, Liu F, & Mao J. (2025). Transcription factor specificity protein (SP) family in renal physiology and diseases. PeerJ, 13, e18820. https://doi.org/10.7717/peerj.18820
[53] Hirohama D, Abedini A, Moon S, Surapaneni A, Dillon ST, Vassalotti A, et al. (2023). Unbiased Human Kidney Tissue Proteomics Identifies Matrix Metalloproteinase 7 as a Kidney Disease Biomarker. J Am Soc Nephrol, 34(7), 1279-1291. https://doi.org/10.1681/ASN.0000000000000141
[54] Tveitaras MK, Skogstrand T, Leh S, Helle F, Iversen BM, Chatziantoniou C, et al. (2015). Matrix Metalloproteinase-2 Knockout and Heterozygote Mice Are Protected from Hydronephrosis and Kidney Fibrosis after Unilateral Ureteral Obstruction. PLoS One, 10(12), e0143390. https://doi.org/10.1371/journal.pone.0143390
[55] Tao Y, Tang C, Wei J, Shan Y, Fang X, & Li Y. (2023). Nr4a1 promotes renal interstitial fibrosis by regulating the p38 MAPK phosphorylation. Mol Med, 29(1), 63. https://doi.org/10.1186/s10020-023-00657-y
[56] Long X, Yan B, & Mo Z. (2023). Uncovering the heterogeneity and cell fate decisions of endothelial cells after myocardial infarction by single‐cell sequencing. Medicine Advances, 1(3), 234-245. https://doi.org/10.1002/med4.34
[57] Dong Z, & Liang X. (2023). A glimpse of recent advances in the research of acute kidney injury. Medicine Advances, 1(2), 158-162. https://doi.org/10.1002/med4.26
Type
Published
Data Availability Statement
The datasets analysed during the current study are available in the GEO Database (GSE66494, https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE66494). In addition, the datasets analyzed during the current study are available from the corresponding author on reasonable request.
Issue
Section
License
Copyright (c) 2026 Life Conflux

This work is licensed under a Creative Commons Attribution 4.0 International License.







