Advancing Precision Medicine through Polygenic Risk Scores: From Statistical Innovation to Clinical Implementation
DOI:
https://doi.org/10.71321/ptbkr794Keywords:
Polygenic risk scores, Genome-wide association studies, Precision medicine, Clinical implementationAbstract
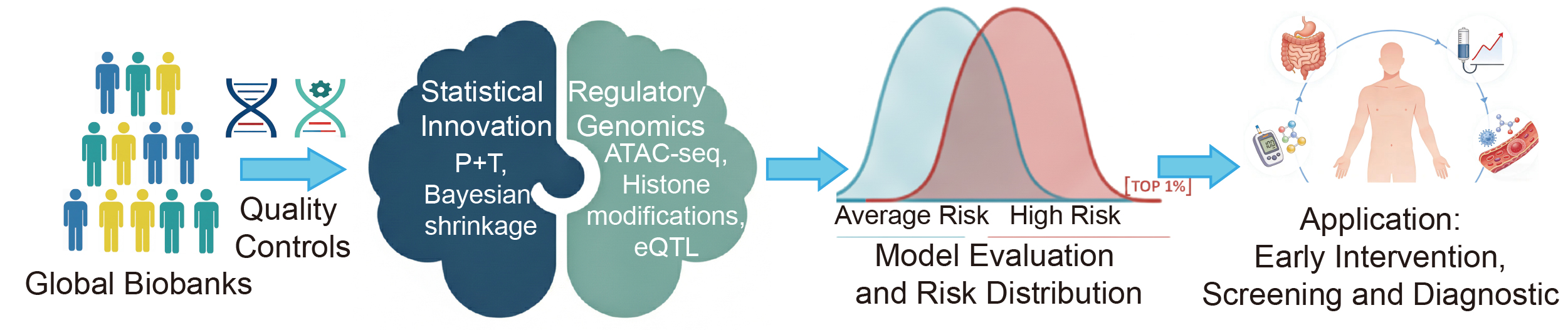
The polygenic risk scores (PRS) have emerged as a transformative approach for quantifying inherited predisposition to complex diseases, leveraging the unprecedented expansion of genome-wide association studies (GWAS) and advances in statistical genetics. By aggregating the marginal effects of millions of common variants, PRS provide a single metric of genetic liability that can achieve predictive performance comparable to traditional clinical risk factors. Current methodologies are undergoing a paradigm shift, moving beyond simple linear additive models to incorporate complex linkage disequilibrium (LD) structures, multi-ancestry frameworks, and functional genomic landscapes. In particular, the integration of regulatory annotations, including expression quantitative trait loci (eQTL), chromatin accessibility, and cell-type-specific enhancers, has enhanced both the biological interpretability and predictive robustness of these scores.
This review synthesizes the rapid methodological evolution of PRS, encompassing Bayesian shrinkage frameworks, machine learning algorithms, and functionally informed strategies designed to mitigate the persistent Eurocentric biases in current datasets. We critically evaluate the evidence supporting the integration of PRS into clinical workflows, focusing on cardiovascular diseases, oncology, and neuropsychiatric disorders, where genetic stratification can enhance preventive interventions and diagnostic precision. Despite this progress, we identify significant challenges to widespread adoption, including the reduced portability of scores across diverse populations, the lack of standardized clinical thresholds, and complex ethical considerations related to health equity.
Finally, we propose a multidisciplinary roadmap for the future of PRS, emphasizing the necessity of global biobank diversity, dynamic risk modeling that incorporates temporal and environmental factors, and the seamless integration of genomic insights into electronic health records. Collectively, these advancements are essential for transitioning PRS from a powerful research tool into an equitable and actionable component of the precision medicine toolkit.
References
[1] Xiang R, Kelemen M, Xu Y, Harris LW, Parkinson H, Inouye M, et al. (2024). Recent advances in polygenic scores: translation, equitability, methods and FAIR tools. Genome Med, 16(1), 33. https://doi.org/10.1186/s13073-024-01304-9
[2] Marcalo R, Neto S, Pinheiro M, Rodrigues AJ, Sousa N, Santos MAS, et al. (2022). Evaluation of the genetic risk for COVID-19 outcomes in COPD and differences among worldwide populations. Plos One, 17(2), e0264009. https://doi.org/10.1371/journal.pone.0264009
[3] Khera AV, Chaffin M, Aragam KG, Haas ME, Roselli C, Choi SH, et al. (2018). Genome-wide polygenic scores for common diseases identify individuals with risk equivalent to monogenic mutations. Nat Genet, 50(9), 1219-1224. https://doi.org/10.1038/s41588-018-0183-z
[4] Mavaddat N, Michailidou K, Dennis J, Lush M, Fachal L, Lee A, et al. (2019). Polygenic Risk Scores for Prediction of Breast Cancer and Breast Cancer Subtypes. American Journal of Human Genetics, 104(1), 21-34. https://doi.org/10.1016/j.ajhg.2018.11.002
[5] Inouye M, Abraham G, Nelson CP, Wood AM, Sweeting MJ, Dudbridge F, et al. (2018). Genomic Risk Prediction of Coronary Artery Disease in 480,000 Adults: Implications for Primary Prevention. J Am Coll Cardiol, 72(16), 1883-1893. https://doi.org/10.1016/j.jacc.2018.07.079
[6] Guseh JS, & Ajufo EC. (2025). Risk in Repose: Sedentary Behavior and Genetic Susceptibility to Cardiovascular Disease. J Am Heart Assoc, 14(6), e041390. https://doi.org/10.1161/JAHA.125.041390
[7] Martin AR, Kanai M, Kamatani Y, Okada Y, Neale BM, & Daly MJ. (2019). Clinical use of current polygenic risk scores may exacerbate health disparities. Nat Genet, 51(4), 584-591. https://doi.org/10.1038/s41588-019-0379-x
[8] Sirugo G, Williams SM, & Tishkoff SA. (2019). The Missing Diversity in Human Genetic Studies. Cell, 177(1), 26-31. https://doi.org/10.1016/j.cell.2019.02.048
[9] Morales J, Welter D, Bowler EH, Cerezo M, Harris LW, McMahon AC, et al. (2018). A standardized framework for representation of ancestry data in genomics studies, with application to the NHGRI-EBI GWAS Catalog. Genome Biol, 19(1), 21. https://doi.org/10.1186/s13059-018-1396-2
[10] Prive F, Arbel J, & Vilhjalmsson BJ. (2021). LDpred2: better, faster, stronger. Bioinformatics, 36(22-23), 5424-5431. https://doi.org/10.1093/bioinformatics/btaa1029
[11] Ge T, Chen CY, Ni Y, Feng YA, & Smoller JW. (2019). Polygenic prediction via Bayesian regression and continuous shrinkage priors. Nature Communications, 10(1), 1776. https://doi.org/10.1038/s41467-019-09718-5
[12] Lloyd-Jones LR, Zeng J, Sidorenko J, Yengo L, Moser G, Kemper KE, et al. (2019). Improved polygenic prediction by Bayesian multiple regression on summary statistics. Nature Communications, 10(1), 5086. https://doi.org/10.1038/s41467-019-12653-0
[13] Skene NG, Bryois J, Bakken TE, Breen G, Crowley JJ, Gaspar HA, et al. (2018). Genetic identification of brain cell types underlying schizophrenia. Nat Genet, 50(6), 825-833. https://doi.org/10.1038/s41588-018-0129-5
[14] Zheng Z, Liu S, Sidorenko J, Wang Y, Lin T, Yengo L, et al. (2024). Leveraging functional genomic annotations and genome coverage to improve polygenic prediction of complex traits within and between ancestries. Nat Genet, 56(5), 767-777. https://doi.org/10.1038/s41588-024-01704-y
[15] Zhao Y, Jhamb D, Shu L, Arneson D, Rajpal DK, & Yang X. (2019). Multi-omics integration reveals molecular networks and regulators of psoriasis. BMC Syst Biol, 13(1), 8. https://doi.org/10.1186/s12918-018-0671-x
[16] Lewis CM, & Vassos E. (2020). Polygenic risk scores: from research tools to clinical instruments. Genome Med, 12(1), 44. https://doi.org/10.1186/s13073-020-00742-5
[17] Torkamani A, Wineinger NE, & Topol EJ. (2018). The personal and clinical utility of polygenic risk scores. Nat Rev Genet, 19(9), 581-590. https://doi.org/10.1038/s41576-018-0018-x
[18] Lambert SA, Gil L, Jupp S, Ritchie SC, Xu Y, Buniello A, et al. (2021). The Polygenic Score Catalog as an open database for reproducibility and systematic evaluation. Nat Genet, 53(4), 420-425. https://doi.org/10.1038/s41588-021-00783-5
[19] Chakshu NK, & Nithiarasu P. (2024). Orbital learning: a novel, actively orchestrated decentralised learning for healthcare. Sci Rep, 14(1), 10459. https://doi.org/10.1038/s41598-024-60915-9
[20] Wang T, Han Z, Yang Y, Tian R, Zhou W, Ren P, et al. (2019). Polygenic Risk Score for Alzheimer's Disease Is Associated With Ch4 Volume in Normal Subjects. Front Genet, 10, 519. https://doi.org/10.3389/fgene.2019.00519
[21] Cope JL, Baukmann HA, Klinger JE, Ravarani CNJ, Bottinger EP, Konigorski S, et al. (2021). Interaction-Based Feature Selection Algorithm Outperforms Polygenic Risk Score in Predicting Parkinson's Disease Status. Front Genet, 12, 744557. https://doi.org/10.3389/fgene.2021.744557
[22] Bycroft C, Freeman C, Petkova D, Band G, Elliott LT, Sharp K, et al. (2018). The UK Biobank resource with deep phenotyping and genomic data. Nature, 562(7726), 203-209. https://doi.org/10.1038/s41586-018-0579-z
[23] Kurki MI, Karjalainen J, Palta P, Sipila TP, Kristiansson K, Donner KM, et al. (2023). FinnGen provides genetic insights from a well-phenotyped isolated population. Nature, 613(7944), 508-518. https://doi.org/10.1038/s41586-022-05473-8
[24] Chen MH, Pitsillides A, & Yang Q. (2021). An evaluation of approaches for rare variant association analyses of binary traits in related samples. Sci Rep, 11(1), 3145. https://doi.org/10.1038/s41598-021-82547-z
[25] Alade A, Mossey P, Awotoye W, Busch T, Oladayo A, Aladenika E, et al. (2024). Rare Variants Analyses Suggest Novel Cleft Genes in the African Population. Res Sq, 10.21203/rs.3.rs-3921355/v1. https://doi.org/10.21203/rs.3.rs-3921355/v1
[26] International Schizophrenia C, Purcell SM, Wray NR, Stone JL, Visscher PM, O'Donovan MC, et al. (2009). Common polygenic variation contributes to risk of schizophrenia and bipolar disorder. Nature, 460(7256), 748-752. https://doi.org/10.1038/nature08185
[27] Visscher PM, Yengo L, Cox NJ, & Wray NR. (2021). Discovery and implications of polygenicity of common diseases. Science, 373(6562), 1468-1473. https://doi.org/10.1126/science.abi8206
[28] Takase M, & Hozawa A. (2025). The Role of Polygenic Risk Score in the General Population: Current Status and Future Prospects. J Atheroscler Thromb, 32(9), 1079-1097. https://doi.org/10.5551/jat.RV22039
[29] Vilhjalmsson BJ, Yang J, Finucane HK, Gusev A, Lindstrom S, Ripke S, et al. (2015). Modeling Linkage Disequilibrium Increases Accuracy of Polygenic Risk Scores. American Journal of Human Genetics, 97(4), 576-592. https://doi.org/10.1016/j.ajhg.2015.09.001
[30] Weissbrod O, Kanai M, Shi H, Gazal S, Peyrot WJ, Khera AV, et al. (2022). Leveraging fine-mapping and multipopulation training data to improve cross-population polygenic risk scores. Nat Genet, 54(4), 450-458. https://doi.org/10.1038/s41588-022-01036-9
[31] Hu Y, Lu Q, Powles R, Yao X, Yang C, Fang F, et al. (2017). Leveraging functional annotations in genetic risk prediction for human complex diseases. PLoS Comput Biol, 13(6), e1005589. https://doi.org/10.1371/journal.pcbi.1005589
[32] Crone B, & Boyle AP. (2024). Enhancing portability of trans-ancestral polygenic risk scores through tissue-specific functional genomic data integration. PLoS Genet, 20(8), e1011356. https://doi.org/10.1371/journal.pgen.1011356
[33] Shieh Y, Hu D, Ma L, Huntsman S, Gard CC, Leung JWT, et al. (2017). Joint relative risks for estrogen receptor-positive breast cancer from a clinical model, polygenic risk score, and sex hormones. Breast Cancer Res Treat, 166(2), 603-612. https://doi.org/10.1007/s10549-017-4430-2
[34] Dimitriou M, Moulos P, Kalafati IP, Saranti G, Rallidis LS, & Dedoussis GV. (2024). Evaluation of Polygenic Risk Scores for Prediction of Coronary Artery Disease in a Greek Case-Control Study. J Pers Med, 14(6). https://doi.org/10.3390/jpm14060565
[35] Guimbaud JB, Siskos AP, Sakhi AK, Heude B, Sabido E, Borras E, et al. (2024). Machine learning-based health environmental-clinical risk scores in European children. Commun Med (Lond), 4(1), 98. https://doi.org/10.1038/s43856-024-00513-y
[36] Carvalho NRG, He Y, Smadbeck P, Flannick J, Mercader JM, Udler M, et al. (2024). Assessing the genetic contribution of cumulative behavioral factors associated with longitudinal type 2 diabetes risk highlights adiposity and the brain-metabolic axis. medRxiv, 10.1101/2024.01.30.24302019. https://doi.org/10.1101/2024.01.30.24302019
[37] He M, Wang D, Xu YJ, Shi J, Liu A, Zhao X, et al. (2025). Multi-omics analysis revealed biomarkers for coronary atherosclerosis: Occurrence and development. Clin Transl Med, 15(8), e70451. https://doi.org/10.1002/ctm2.70451
[38] Tang H, & He Z. (2021). Advances and challenges in quantitative delineation of the genetic architecture of complex traits. Quant Biol, 9(2), 168-184. https://doi.org/10.15302/j-qb-021-0249
[39] Vassy JL, Durant NH, Kabagambe EK, Carnethon MR, Rasmussen-Torvik LJ, Fornage M, et al. (2012). A genotype risk score predicts type 2 diabetes from young adulthood: the CARDIA study. Diabetologia, 55(10), 2604-2612. https://doi.org/10.1007/s00125-012-2637-7
[40] Natarajan P, Young R, Stitziel NO, Padmanabhan S, Baber U, Mehran R, et al. (2017). Polygenic Risk Score Identifies Subgroup With Higher Burden of Atherosclerosis and Greater Relative Benefit From Statin Therapy in the Primary Prevention Setting. Circulation, 135(22), 2091-2101. https://doi.org/10.1161/CIRCULATIONAHA.116.024436
[41] Stein R, Ferrari F, & Garcia-Giustiniani D. (2024). Polygenic Risk Scores: The Next Step for Improved Risk Stratification in Coronary Artery Disease? Arq Bras Cardiol, 121(9), e20240252. https://doi.org/10.36660/abc.20240252 (Escores de Risco Poligenico: O Proximo Passo para Melhorar a Estratificacao de Risco na Doenca Arterial Coronariana?)
[42] Seibert TM, Fan CC, Wang Y, Zuber V, Karunamuni R, Parsons JK, et al. (2018). Polygenic hazard score to guide screening for aggressive prostate cancer: development and validation in large scale cohorts. BMJ, 360, j5757. https://doi.org/10.1136/bmj.j5757
[43] Greenberg R, Aharonov-Majar E, Isakov O, Hayek S, Elefant N, Balicer RD, et al. (2023). Carrier screening program for BRCA1/BRCA2 pathogenic variants among Ashkenazi Jewish women in Israel: An observational study. Genet Med Open, 1(1), 100824. https://doi.org/10.1016/j.gimo.2023.100824
[44] Hassanin E, Maj C, Klinkhammer H, Krawitz P, May P, & Bobbili DR. (2023). Assessing the performance of European-derived cardiometabolic polygenic risk scores in South-Asians and their interplay with family history. BMC Med Genomics, 16(1), 164. https://doi.org/10.1186/s12920-023-01598-5
[45] Udler MS, McCarthy MI, Florez JC, & Mahajan A. (2019). Genetic Risk Scores for Diabetes Diagnosis and Precision Medicine. Endocr Rev, 40(6), 1500-1520. https://doi.org/10.1210/er.2019-00088
[46] Makowski C, Wang H, & Chen CH. (2022). Clinical opportunity awaits at the intersection of genomics and brain imaging. J Psychiatry Neurosci, 47(4), E293-E298. https://doi.org/10.1503/jpn.220075
[47] Du Y, Hao H, Wang S, Pearlson GD, & Calhoun VD. (2020). Identifying commonality and specificity across psychosis sub-groups via classification based on features from dynamic connectivity analysis. Neuroimage Clin, 27, 102284. https://doi.org/10.1016/j.nicl.2020.102284
[48] Motazedi E, Cheng W, Thomassen JQ, Frei O, Rongve A, Athanasiu L, et al. (2022). Using Polygenic Hazard Scores to Predict Age at Onset of Alzheimer's Disease in Nordic Populations. Journal of Alzheimers Disease, 88(4), 1533-1544. https://doi.org/10.3233/JAD-220174
[49] Desikan RS, Fan CC, Wang Y, Schork AJ, Cabral HJ, Cupples LA, et al. (2017). Genetic assessment of age-associated Alzheimer disease risk: Development and validation of a polygenic hazard score. PLoS Med, 14(3), e1002258. https://doi.org/10.1371/journal.pmed.1002258
[50] Tan CH, Bonham LW, Fan CC, Mormino EC, Sugrue LP, Broce IJ, et al. (2019). Polygenic hazard score, amyloid deposition and Alzheimer's neurodegeneration. Brain, 142(2), 460-470. https://doi.org/10.1093/brain/awy327
[51] Singh S, Stocco G, Theken KN, Dickson A, Feng Q, Karnes JH, et al. (2024). Pharmacogenomics polygenic risk score: Ready or not for prime time? Clin Transl Sci, 17(8), e13893. https://doi.org/10.1111/cts.13893
[52] Zheng SL, Jurgens SJ, McGurk KA, Xu X, Grace C, Theotokis PI, et al. (2025). Evaluation of polygenic scores for hypertrophic cardiomyopathy in the general population and across clinical settings. Nat Genet, 57(3), 563-571. https://doi.org/10.1038/s41588-025-02094-5
[53] Ruan Y, Lin YF, Feng YA, Chen CY, Lam M, Guo Z, et al. (2022). Improving polygenic prediction in ancestrally diverse populations. Nat Genet, 54(5), 573-580. https://doi.org/10.1038/s41588-022-01054-7
[54] Harley ITW, & Sawalha AH. (2022). Systemic lupus erythematosus as a genetic disease. Clin Immunol, 236, 108953. https://doi.org/10.1016/j.clim.2022.108953
[55] Bray NJ, & O'Donovan MC. (2019). The genetics of neuropsychiatric disorders. Brain Neurosci Adv, 2. https://doi.org/10.1177/2398212818799271
[56] Wray NR, Lee SH, Mehta D, Vinkhuyzen AA, Dudbridge F, & Middeldorp CM. (2014). Research review: Polygenic methods and their application to psychiatric traits. J Child Psychol Psychiatry, 55(10), 1068-1087. https://doi.org/10.1111/jcpp.12295
[57] Choi SW, Mak TS, & O'Reilly PF. (2020). Tutorial: a guide to performing polygenic risk score analyses. Nat Protoc, 15(9), 2759-2772. https://doi.org/10.1038/s41596-020-0353-1
[58] Abou Tayoun A, Ali H, & Mokrab Y. (2025). The quest for a complete understanding of the human genome. Genome Med, 17(1), 88. https://doi.org/10.1186/s13073-025-01523-8
[59] Ovretveit K, Ingestrom EML, Spitieris M, Tragante V, Wade KH, Thomas LF, et al. (2024). Polygenic risk scores associate with blood pressure traits across the lifespan. Eur J Prev Cardiol, 31(6), 644-654. https://doi.org/10.1093/eurjpc/zwad365
[60] Bierut LJ. (2019). Using genomic profiling for understanding and improving response to smoking cessation treatment. Curr Epidemiol Rep, 6(4), 486-490. https://doi.org/10.1007/s40471-019-00220-6
[61] Gurdasani D, Barroso I, Zeggini E, & Sandhu MS. (2019). Genomics of disease risk in globally diverse populations. Nat Rev Genet, 20(9), 520-535. https://doi.org/10.1038/s41576-019-0144-0
[62] Consortium HA, Rotimi C, Abayomi A, Abimiku A, Adabayeri VM, Adebamowo C, et al. (2014). Research capacity. Enabling the genomic revolution in Africa. Science, 344(6190), 1346-1348. https://doi.org/10.1126/science.1251546
[63] Gao M, Zheng Q, Jiang Y, Chang X, & Zheng X. (2025). Evaluation of diverse polygenic risk score models for lung cancer in a small-scale Chinese cohort. Front Genet, 16, 1646997. https://doi.org/10.3389/fgene.2025.1646997
[64] Bryois J, Skene NG, Hansen TF, Kogelman LJA, Watson HJ, Liu Z, et al. (2020). Genetic identification of cell types underlying brain complex traits yields insights into the etiology of Parkinson's disease. Nat Genet, 52(5), 482-493. https://doi.org/10.1038/s41588-020-0610-9
[65] Dong S, & Boyle AP. (2022). Prioritization of regulatory variants with tissue-specific function in the non-coding regions of human genome. Nucleic Acids Research, 50(1), e6. https://doi.org/10.1093/nar/gkab924
[66] Smelik M, Zhao Y, Li X, Loscalzo J, Sysoev O, Mahmud F, et al. (2024). An interactive atlas of genomic, proteomic, and metabolomic biomarkers promotes the potential of proteins to predict complex diseases. Sci Rep, 14(1), 12710. https://doi.org/10.1038/s41598-024-63399-9
[67] Kelemen M, Xu Y, Jiang T, Zhao JH, Anderson CA, Wallace C, et al. (2025). Performance of deep-learning-based approaches to improve polygenic scores. Nature Communications, 16(1), 5122. https://doi.org/10.1038/s41467-025-60056-1
[68] Schuran M, Goudey B, Dite GS, & Makalic E. (2025). A survey on deep learning for polygenic risk scores. Brief Bioinform, 26(4). https://doi.org/10.1093/bib/bbaf373
[69] Pigoni A, Delvecchio G, Turtulici N, Madonna D, Pietrini P, Cecchetti L, et al. (2024). Machine learning and the prediction of suicide in psychiatric populations: a systematic review. Transl Psychiatry, 14(1), 140. https://doi.org/10.1038/s41398-024-02852-9
[70] Harrison H, Li N, Saunders CL, Rossi SH, Dennis J, Griffin SJ, et al. (2022). The current state of genetic risk models for the development of kidney cancer: a review and validation. BJU Int, 130(5), 550-561. https://doi.org/10.1111/bju.15752
[71] Carmienke S, Baumert J, Gabrys L, Heise M, Frese T, Heidemann C, et al. (2020). Participation in structured diabetes mellitus self-management education program and association with lifestyle behavior: results from a population-based study. BMJ Open Diabetes Res Care, 8(1). https://doi.org/10.1136/bmjdrc-2019-001066
[72] Lerga-Jaso J, Terpolovsky A, Novkovic B, Osama A, Manson C, Bohn S, et al. (2025). Optimization of multi-ancestry polygenic risk score disease prediction models. Sci Rep, 15(1), 17495. https://doi.org/10.1038/s41598-025-02903-1
[73] Kember RL, Merikangas AK, Verma SS, Verma A, Judy R, Regeneron Genetics C, et al. (2021). Polygenic Risk of Psychiatric Disorders Exhibits Cross-trait Associations in Electronic Health Record Data From European Ancestry Individuals. Biol Psychiatry, 89(3), 236-245. https://doi.org/10.1016/j.biopsych.2020.06.026
[74] Visscher PM, Wray NR, Zhang Q, Sklar P, McCarthy MI, Brown MA, et al. (2017). 10 Years of GWAS Discovery: Biology, Function, and Translation. American Journal of Human Genetics, 101(1), 5-22. https://doi.org/10.1016/j.ajhg.2017.06.005
[75] Wang L, Khunsriraksakul C, Markus H, Chen D, Zhang F, Chen F, et al. (2024). Integrating single cell expression quantitative trait loci summary statistics to understand complex trait risk genes. Nature Communications, 15(1), 4260. https://doi.org/10.1038/s41467-024-48143-1
[76] Visvikis-Siest S, Stathopoulou MG, Sunder-Plassmann R, Alizadeh BZ, Barouki R, Chatzaki E, et al. (2023). The 10th Santorini conference: Systems medicine, personalised health and therapy. "The odyssey from hope to practice: Patient first. Keep Ithaca always in your mind", Santorini, Greece, 23-26 May 2022. Front Genet, 14, 1171131. https://doi.org/10.3389/fgene.2023.1171131
Type
Published
Data Availability Statement
Not Applicable
Issue
Section
License
Copyright (c) 2026 Brain Conflux

This work is licensed under a Creative Commons Attribution 4.0 International License.







